Jasmine Otto is a visualization developer who designs novel intelligent tools and interfaces, by working closely with domain experts. Her work is published in selective venues including IEEE VIS and AAAI AIIDE. She holds a PhD in Computational Media from the Design Reasoning Lab at UC Santa Cruz, and has worked for NASA JPL on robotics operations scheduling.
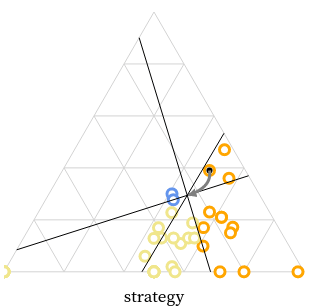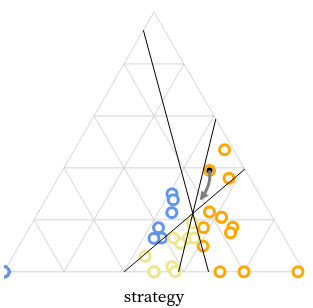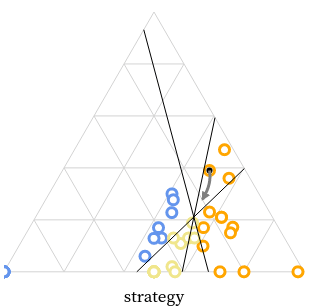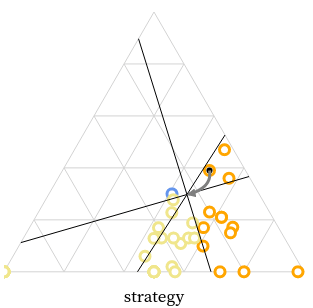
We created the MarsIPAN (Mars Interactive Pass Allocation Navigator) schedule, and a companion dashboard for comparing allocations. Operations planners are using these visualizations to assess both operational efficiency and staffing requirements across many possible scenarios.
See also our Data to Discovery 2021 project page. With thanks to the DTD team and our JPL stakeholders.